-Search query
-Search result
Showing 1 - 50 of 1,958 items for (author: feng & q)

EMDB-36849: 
Nipah virus Attachment glycoprotein with 41-6 antibody fragment
Method: single particle / : Sun MM

PDB-8k3c: 
Nipah virus Attachment glycoprotein with 41-6 antibody fragment
Method: single particle / : Sun MM

EMDB-43991: 
Cryo-EM structure of apo state human Cav3.2
Method: single particle / : Fan X, Huang J, Yan N

EMDB-43992: 
Cryo-EM structure of human Cav3.2 with TTA-A2
Method: single particle / : Fan X, Huang J, Yan N

EMDB-43993: 
Cryo-EM structure of human Cav3.2 with TTA-P2
Method: single particle / : Fan X, Huang J, Yan N

EMDB-43994: 
Cryo-EM structure of human Cav3.2 with ML218
Method: single particle / : Fan X, Huang J, Yan N

EMDB-43995: 
Cryo-EM structure of human Cav3.2 with ACT-709478
Method: single particle / : Fan X, Huang J, Yan N

PDB-9ayg: 
Cryo-EM structure of apo state human Cav3.2
Method: single particle / : Fan X, Huang J, Yan N

PDB-9ayh: 
Cryo-EM structure of human Cav3.2 with TTA-A2
Method: single particle / : Fan X, Huang J, Yan N

PDB-9ayj: 
Cryo-EM structure of human Cav3.2 with TTA-P2
Method: single particle / : Fan X, Huang J, Yan N

PDB-9ayk: 
Cryo-EM structure of human Cav3.2 with ML218
Method: single particle / : Fan X, Huang J, Yan N

PDB-9ayl: 
Cryo-EM structure of human Cav3.2 with ACT-709478
Method: single particle / : Fan X, Huang J, Yan N
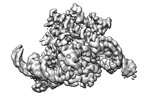
EMDB-35323: 
Cryo-EM structure of the ISFba1 TnpB-reRNA-dsDNA complex
Method: single particle / : Yin M, Zhou F, Zhu Y, Huang Z

PDB-8iaz: 
Cryo-EM structure of the ISFba1 TnpB-reRNA-dsDNA complex
Method: single particle / : Yin M, Zhou F, Zhu Y, Huang Z

EMDB-37154: 
Cyanophage A-1(L) neck/gp7-terminator
Method: single particle / : Yu RC, Li Q, Zhou CZ

EMDB-37155: 
Cyanophage A-1(L) neck/gp5-neck fiber
Method: single particle / : Yu RC, Li Q, Zhou CZ

EMDB-37150: 
The CBD domain of cyanophage A-1(L) short tail fiber
Method: single particle / : Yu RC, Li Q, Zhou CZ

EMDB-37151: 
Cyanophage A-1(L) baseplate-initiators
Method: single particle / : Yu RC, Li Q, Zhou CZ

EMDB-37152: 
Cyanophage A-1(L) tail fiber
Method: single particle / : Yu RC, Li Q, Zhou CZ

EMDB-37153: 
Cyanophage A-1(L) sheath-tube
Method: single particle / : Yu RC, Li Q, Zhou CZ

EMDB-41590: 
Cyanophage A-1(L) portal
Method: single particle / : Yu RC, Li Q, Zhou CZ

PDB-8ke9: 
The CBD domain of cyanophage A-1(L) short tail fiber
Method: single particle / : Yu RC, Li Q, Zhou CZ

EMDB-37104: 
96-nm axonemal repeat with RS1/2/3
Method: subtomogram averaging / : Cong X, Yao C

EMDB-37111: 
48-nm repeat DMT
Method: subtomogram averaging / : Cong X, Yao C

EMDB-37114: 
Radial Spoke 1 (RS1)
Method: subtomogram averaging / : Cong X, Yao C

EMDB-37116: 
RS1 refined with head mask
Method: subtomogram averaging / : Cong X, Yao C

EMDB-37117: 
Radial Spoke 2 (RS2)
Method: subtomogram averaging / : Cong X, Yao C

EMDB-37118: 
Radial Spoke 2 (RS2) head
Method: subtomogram averaging / : Cong X, Yao C

EMDB-37119: 
Radial Spoke 3
Method: subtomogram averaging / : Cong X, Yao C

EMDB-37120: 
Radial Spoke 3 head
Method: subtomogram averaging / : Cong X, Yao C

EMDB-34848: 
Structure of PKD2-F604P (Polycystin-2, TRPP2) with ML-SA1
Method: single particle / : Chen MY, Su Q, Wang ZF, Yu Y

PDB-8hk7: 
Structure of PKD2-F604P (Polycystin-2, TRPP2) with ML-SA1
Method: single particle / : Chen MY, Su Q, Wang ZF, Yu Y

EMDB-18267: 
Structure of the human 20S U5 snRNP core
Method: single particle / : Schneider S, Galej WP

EMDB-19041: 
Structure of the human 20S U5 snRNP
Method: single particle / : Schneider S, Galej WP

PDB-8q91: 
Structure of the human 20S U5 snRNP core
Method: single particle / : Schneider S, Galej WP

PDB-8rc0: 
Structure of the human 20S U5 snRNP
Method: single particle / : Schneider S, Galej WP

EMDB-36301: 
Cryo-EM structure of the TcsH-TMPRSS2 complex
Method: single particle / : Zhou R, Tao L, Zhan X

EMDB-36302: 
The cryo-EM map of the C-terminal region in the TcsH-TMPRSS2 complex
Method: single particle / : Zhou R, Tao L, Zhan X

EMDB-36303: 
Cryo-EM structure of the TcsH-CROP in complex with TMPRSS2
Method: single particle / : Zhou R, Tao L, Zhan X

PDB-8jhz: 
Cryo-EM structure of the TcsH-TMPRSS2 complex
Method: single particle / : Zhou R, Liang T, Zhan X

PDB-8ji0: 
Cryo-EM structure of the TcsH-CROP in complex with TMPRSS2
Method: single particle / : Zhou R, Tao L, Zhan X
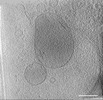
EMDB-39012: 
Representative tomogram of primary glioblastoma stem cell with circular inter-mitochondrial junctions.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT

EMDB-39015: 
Representative tomogram of microglia cell with nanotunnel-like structures resembling mitochondrial fission.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT
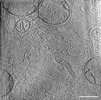
EMDB-39019: 
Representative tomogram of glioblastoma cell with nanotunnel-like structure and inter-mitochondrial junction.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search









 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
